NanoCommons Demonstration Case - Best-practice in SOP development for nanosafety assessment
Currently, the limited sharing and the free-text format of standard operating procedures makes independent quality control and comparison of different SOPs almost impossible. Therefore, this demonstration case is meant to document the SOP development process and derive a checklist for assessment of SOP quality and completeness based on standardised reporting formats. This case is ongoing.
Table of contents
Background
Performing experiments following standard operating procedures (SOPs) is essential in industrial settings to guarantee reproducibility and in regulatory settings for validation of the method and acceptance of the result and conclusions. SOPs are now also more and more implemented in basic- and early-research academic and industrial laboratories. Developing these SOPs is a complex and time-consuming process but it is rarely documented with respect to the thought processes, which went into the SOP, and quality assurance measures adopted during the development. Additionally, SOPs are in most cases only shared if absolutely needed e.g., for regulatory validation and even if they are, the free-text format makes independent quality control and comparison of different SOPs almost impossible. Taking all of this together, there are clear gaps needed to be filled before general guidelines for SOP development and quality and completeness evaluations can be proposed and used for training of early-stage researchers. Therefore, this demonstration case is meant to document the SOP development process and derive a checklist for assessment of SOP quality and completeness. The basis of the case study will be the well documented SOPs for assessment for cell seeding and cell viability (MTS assay for cytotoxicity) developed and round robin tested in the FP7 project QualityNano, including extensive evaluation of the discrepancies in the data produced by the partners, then further refined and complemented with a full set of training materials before being successfully utilised in the FP7 project NeuroNano. The latter project included the partners who had developed the SOP and training materials and several new partners who had not previously participated in the SOP development or training. Key aspects of the SOP developed and optimisation included evaluation of user proficiency before running the SOP, training of the participants (via webinar) to address some of the key areas where errors can occur, and collection of detailed metadata regarding the participants experience of implementing the SOP.
The post assay completion survey, to be completed by all partners once they had undertaken the assay following the SOP, was used to assess their compliance with the subtle details of the protocol and to aid in the analysis of results, specifically in regards to any anomalous or deviant results. A snapshot is shown in Figure 1.
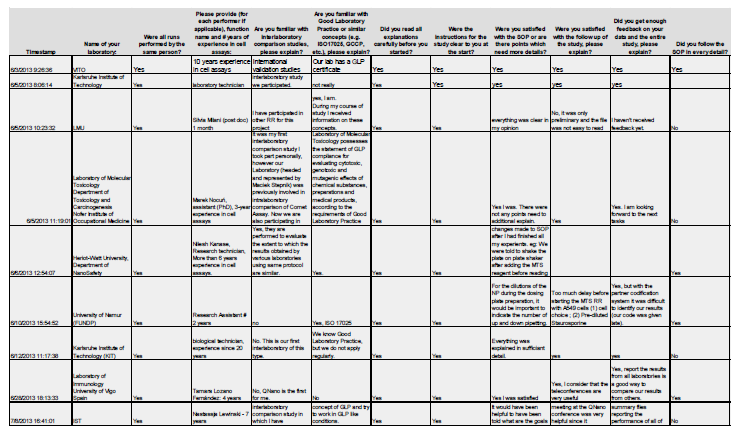 Figure 1. Snapshot of results from the QualityNano Questionnaire on performance of the MTS assay SOP (to be filled out by the person performing the assay in order to help the lead partner identify any potential issues).
Figure 1. Snapshot of results from the QualityNano Questionnaire on performance of the MTS assay SOP (to be filled out by the person performing the assay in order to help the lead partner identify any potential issues).
By re-assessing the approaches, documentation and training developed in QualityNano, and its successful deployment in FutureNanoNeeds, this case study will attempt to extract out some key steps and lessons for the development, documentation and systematization of knowledge sharing related to nanosafety best practice. Figure 2 shows the multiple steps in the MTS assay and the critical steps that were identified in implementing the SOP.
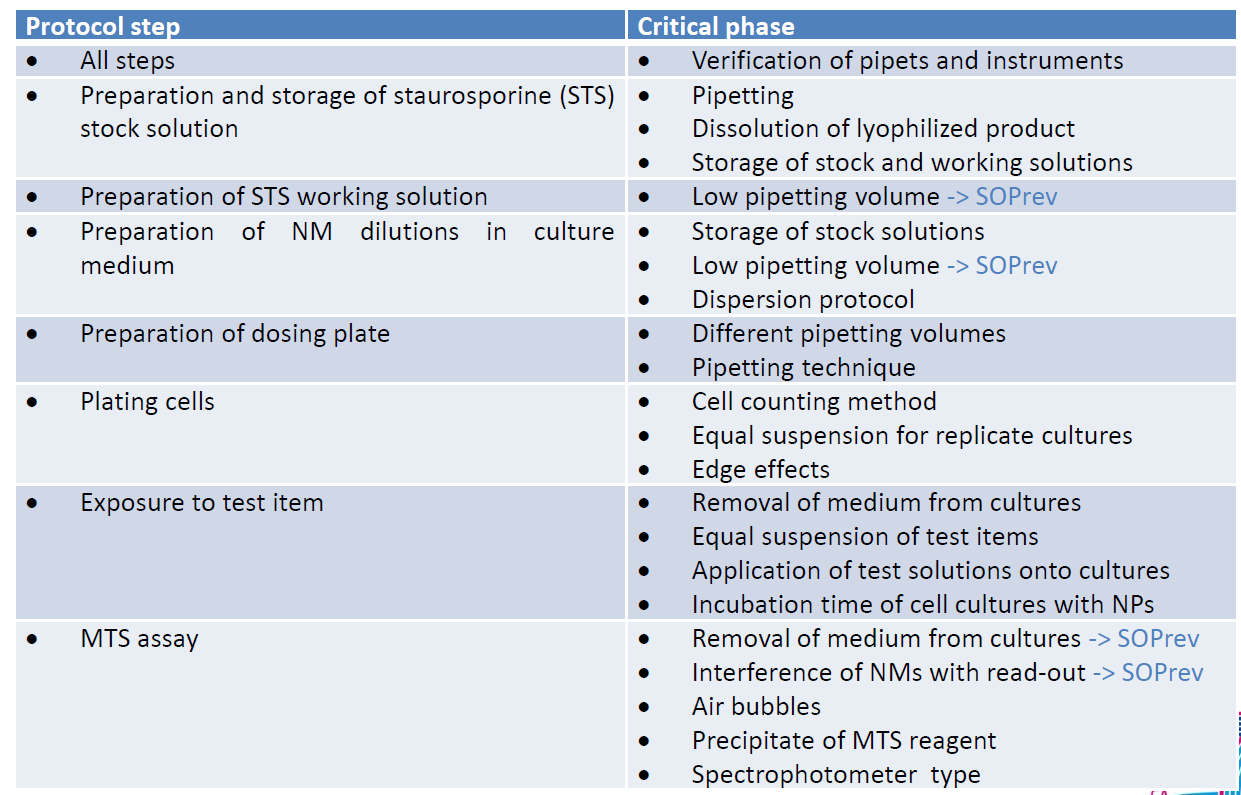 Figure 2. Protocol steps in the MTS assay and the critical phases that were identified in implementing the SOP. Note that many of these, such as pipetting technique and edge effects, are usually assumed knowledge passed on from lab-member to lab-member rather than being documented in experimental methods sections in publications, or even documented in protocols. Thus, highlighting these deserves action. For instance, this also includes important visual checks ensuring equal volumes for replicate cultures (which again seems obvious but with multi-channel pipettes needs checking).
Figure 2. Protocol steps in the MTS assay and the critical phases that were identified in implementing the SOP. Note that many of these, such as pipetting technique and edge effects, are usually assumed knowledge passed on from lab-member to lab-member rather than being documented in experimental methods sections in publications, or even documented in protocols. Thus, highlighting these deserves action. For instance, this also includes important visual checks ensuring equal volumes for replicate cultures (which again seems obvious but with multi-channel pipettes needs checking).
We will also review the ACEnano deliverables from its WP5 on “SOPs and method validation” to see what can be systematized there. These will then be used, on the one hand, to start a best-practice guideline and, on the other hand, to evaluate existing tools within NanoCommons and neighbouring projects on their usability to support more structured SOP development, to design and start developing new services supporting this and to showcase the benefits of a structured and annotated repository of nanosafety SOPs for direct use in other laboratories. This will in turn improve reproducibility and facilitate adapting it to other techniques and for training purposes. Given that much of this applies beyond nanosafety, the translatability to other research communities will be high, but we will focus on aspects that are particular to nanomaterials (e.g., their dynamic nature, their surface interactions etc.) and those specific to the toxicology / ecotoxicology communities in particular, leveraging also the expertise in the wider ELIXIR toxicology community that NanoCommons has been a founding member of.
Aims
This demonstration case will aim to develop a systematized process for SOP development to capture the inherent knowledge that is often not fully documented but is key to reproducibility and best practice. Based on these guidelines, we will evaluate if reporting standards and templates can be developed to harmonise the collection of such knowledge, make it better comparable to see differences in similar SOPs, e.g., for the same endpoint and identify optimisation options for a specific study, and perhaps could even provide structured metadata to be provided as part of datasets performed according to this SOP. This could be aligned with concepts developed for the MODA and CHADA templates (see below), which provide not only information on the experimental procedure but also on the measurement principle, the theoretical background and applicability domain.
Such structured reporting could also help to improve the reusability of the description, e.g., as part of a complex experimental setup, where existing SOPs could be used as part of the overall description by linking in the structured reports. The NIKC concept of instances described above for the study design reporting could also be coming in handy here to give a visual representation of the linked SOPs and their relationships to each other.
Tasks
This demonstration case will evaluate the experiences from QualityNano and FutureNanoNeeds on development and optimisation of the SOP for assessment of NMs cytotoxicity by MTS assay, and utilise this to develop a set of guidelines and best practice examples depicting how to document SOPs in sufficient detail to maximise reproducibility and facilitate complete metadata capture to support data re-usability.
Evaluate the ACEnano guidance documents on SOP development and method validation to develop a checklist of features that must be documented as part of an SOP for toxicity evaluation as a basis to develop a structured SOP development workflow. This will also include annotation of the documents with relevant ontology terms from the eNanoMapper (ENM) ontology and where no suitable terms are identified use of the NSC Terminology Harmoniser to add the terms and have them approved for inclusion in the ENM ontology.
Utilising the collected best practice and the checklist, a set of 10 biological assays spanning areas where the NanoCommons partners have expertise (e.g., immunotoxicity, protein corona determination, acute and chronic daphnia toxicity, ecotoxicity to worms, cytotoxicity including confirmation of NMs uptake and localization, etc.) will be developed, ontologically annotated, and structured in a coherent and systematic way. This can also be supported by efforts underway in the RiskGONE project to revise the chronic reproductive assay for daphnia, and the genotoxicity testing in fish cells lines and whole daphnia. These will be integrated into protocol.io and used as the basis of an SOP-generator (equivalent to the MODA and QMRF generators, or the DMP tools currently available) via a guided web-form linked to the checklist, and providing guidance on what needs to be included, how to identify the appropriate ontological terms, considerations in terms of calibration, visual inspection, assumed knowledge etc. Walking that process will also form the metadata needed to enable re-use of data generated utilising the protocols.
Using the exemplar structures guided by SOPs, the final activity will be to showcase the benefits of a structured and annotated repository of nanosafety SOPs for direct use in other laboratories to improve reproducibility, for adaptation to other techniques and for training purposes. This will include demonstrations in RiskGONE, and support for the wider set of SOPs being developed there on human toxicity and NMs characterisation, and to the S(S)bD and other project clusters.
Expected outcomes
With this demonstration case we expect to evaluate what should be in an SOP in order to provide quality-assured outcome of a set of 10 biological assays, i.e. to be complete for purpose in metadata collection, which means in respect to the intended biological endpoints, and to ensure sufficient detail on how to generate the data to maximise reproducibility. Answers to the following questions shall be obtained:
- Can we structure the SOP in a way to get structured metadata out?
- How does it relate to other aspects - and is it possible to integrate the whole workflow from experimental design guided by an SOP-type step-by-step protocol through to the datasets generated.
- What is the overlap in specific examples taken from this demonstration case on SOPs with examples of biological assay data templates designed in Demonstration Case - Best-practice in nanosafety study design and its documentation via visual experimental maps?
The above-described cross-evaluation of SOPs with biological assays data templates for upload into the NanoCommons KB will also include the integration of SOPs available from Demonstration Case Electronic Laboratory Notebooks for data collection and annotation and Demonstration Case Alignment of in silico protein corona modelling with experimental analysis.