Integration of public data warehouses
The harmonisation and integration of these existing data infrastructures from completed or ongoing projects, along with potential expansion to incorporate advanced materials in general, requires certain workflows and processes to be in place.
These are currently set up by the NanoCommons project to facilitate the operational procedure for database integration (and dataset upload for isolated datasets as described in the Generation and upload of individual datasets, and to provide end users querying the NanoCommons Knowledge Base (KB) with the best possible outcome integrating datasets pulled from the widest range of federated and semantically mapped databases possible. Such workflows require the exchange of information between the harmonisation layer of the NanoCommons KB and the source databases like the NanoCommons Data Warehouse (DW), which includes:
- Specific details of both project manager and technical personnel that will be able to address or redirect any identified issues;
- the used operating system, software and APIs;
- a detailed mapping of the database hierarchical structure;
- the data types (NM and media chemistry characterisation, linked to NM properties (intrinsic and extrinsic) and hazard endpoints, across exposure time(series), details of methods, organisms and cells utilised etc.), size and formats;
- the enrichment processes (if any) that have been utilised or that are required;
- the applied quality checks (e.g. data cleansing, completeness, competence or certification level of the curator);
- the metadata requirements and availability for each data type (see iv. above);
- the presence or not of DOIs and/or unique database IDs for the overall database or subsets of the data within the database;
- data annotation – which ontology(ies) has/have been used;
- the applied measures and validation and reconciliation processes to prevent external security breaches;
- error handling processes;
- information on whether the database is complete (containing data from finalised projects) or if active how frequently it is being updated and the volumes of data added in each update; and
- information on the long-term sustainability / hosting plan for the database in its current location and the current duration of the hosting arrangements.
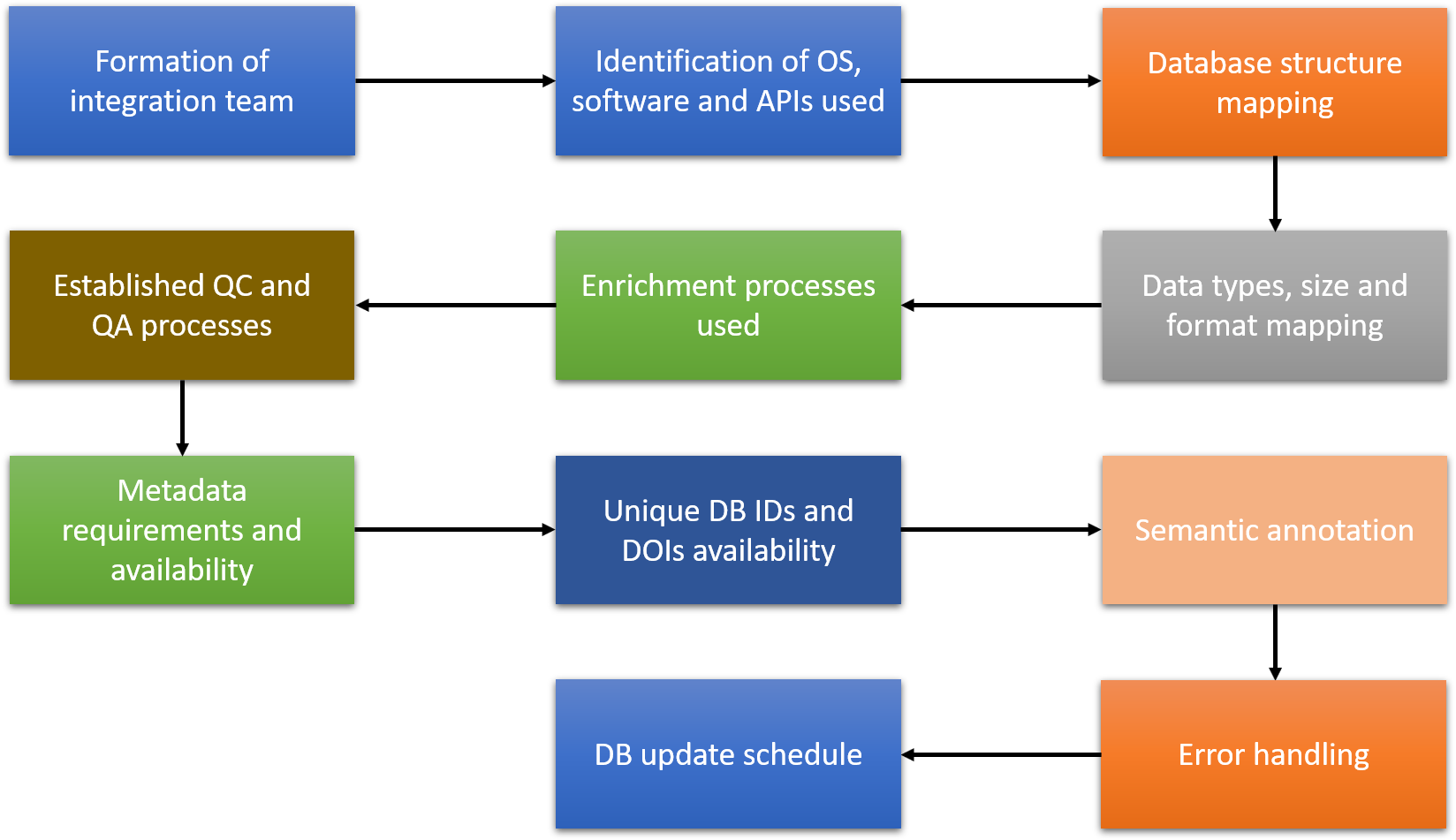 Figure 1. NanoCommons workflow for database integration into the KB.
Figure 1. NanoCommons workflow for database integration into the KB.
Based on this information, the workflow shown in Figure 1 will be worked through with the existing database provider, with re-interaction with data owner/generator for information gathering steps as needed to ensure all required information is available. Once the information has been collected, the NanoCommons database integration team (made up of NanoCommons technical expert representatives (Biomax, Seven Past Nine, UoB) and the key representatives of the source database) will apply the necessary semantic mapping, and test the communication between the NanoCommons KB and the integrated or federated database. Once the testing has been completed, an announcement will be made via the NanoCommons website and via the NSC newsletter and other relevant dissemination channels. In the same way as for the integration of complete databases, a dedicated workflow is followed to collect the information required to facilitate incorporation of individual datasets into the NanoCommons KB. Most of the steps followed are identical or aligned between the two ways of data integration. These include the information gathering steps needed to ensure all required information is available to host the data in the NanoCommons KB, application of the necessary annotation of the dataset and the semantic mapping, and assessment of the data completeness and quality to tag in the metadata for feedback to users querying the datasets. A complete description of each step in the database and dataset integration into NanoCommons is presented in the next sections, and the full checklist for source DBs is included in the Appendix.
Database integration procedure
NanoCommons follows the concept of database integration to prevent unnecessary duplication of large datasets with the associated energy costs, with the exception of small datasets or ended projects whose data is not available anywhere else (mode 2 indicated above). This means that the data is not extracted from the original databases and stored in a common data warehouse, but becomes a “federated” dataset where the data stays in the original place and is linked with other data sources via the semantic layer for harmonised, interoperable access from the NanoCommons research infrastructure platform. The correct integration of remote databases is, therefore, one of the most significant processes of the NanoCommons e-infrastructures in order to be able identify (findable), share (accessible) and reuse (interoperable and reusable) information existing in different sources. In order for the integration to be successful, the detailed guidance for the workflow described above needs to be established that will facilitate the exchange of summary and raw data with the corresponding metadata, based on well-defined steps that will be followed by experts at both the source database, the NanoCommons KB and users (individuals and/or source databasess). These steps can be sorted into 5 separate categories, each one containing specific functionalities:
- Communication, monitoring, management and support
- Connectivity, delivery and deployment
- Data transformation
- Data governance and security to ensure long-term sustainability of the integrated database
- Tools integration.
These five steps are described in further detail below, and are mapped into the workflows show in Figure 1.
Communication, monitoring, management and support
Communication
The first step in database integration is to establish a clear and direct communication between the expert personnel of the NC consortium and the respective experts/developers on the side of the database to be integrated (the source DB). The identified experts will be responsible for exchanging the necessary information to achieve DB connectivity and harmonisation and will form the integration team for the specific DB. This team will be responsible, besides the technical issues, to align and harmonise the semantic annotation of the data model (data schema) of the source DB with NanoCommons as well as the data and metadata to ensure maximum compatibility and interoperability. Especially, it is essential that the data schema and the NM description use the same terminology (ontology) since they are used for linking equivalent data entries in different databases and allow search and browsing of NMs across databases, respectively. In this step, a comprehensive agreement between the NanoCommons project and the source DB will also be formed covering aspects such as the frequency of updates of the database, and the long-term maintenance and quality assurance processes for the source DB (as per the outline shown in the Appendix).
Monitoring and management
One of the key practices of DB management and data integration is the continuous monitoring of the integration process to make sure that any exceptions and glitches reported will be handled and resolved promptly. This will be achieved through a centralised and web-based monitoring system. As a result, source DB owners that would like to be integrated into the KB will need to provide specific information on the known issues, exceptions and limitations existing in their DBs, significant updates that may affect functionality and provide proper notifications on scheduled maintenance work and offline time in general.
Support
The source DB owners will need to provide the NC expert team with detailed information on the support practices available through their team and how they plan to resolve any issues that may arise during and after integration. The information will be evaluated by the NC consortium and specific guidelines will be developed. Additional training might be requested in terms of the source DB functionality.
Connectivity, delivery and deployment
The continuous connectivity and communication between the source DB and the NanoCommons KB are imperative for the successful integration and data implementation into the NanoCommons KB. In that sense, the following information needs to be provided to the NanoCommons experts.
Source database Operating System, size and mapping
To establish proper communication and sourcing from remote databases, information on the operation system (OS) used to build the DB is needed in order to test compatibility with the NanoCommons KB and to identify any necessary modifications/adaptations needed to allow communication. Information will also be needed on the load capacity of the DB and its ability to handle increased traffic, especially during the use of tools implemented in the NanoCommons KB. At the same time, a complete mapping of the source DB hierarchical structure and the total size of the various types of data included and metadata will be needed. The source DB owner will also need to provide information on the protocols supported from the DB for data movement, sharing and synchronisation. Such information would be used to achieve maximum synchronisation between the databases and/or KB and achieve maximum up time, i.e. time when the system is working and NanoCommons users are able to seamlessly access federated data.
Data transformation
All of the processes used to curate, structure, annotate and enrich data and metadata in the source database fall under this category. The information to be asked for will be whether the database uses a structured and harmonised curation template for data entry. The existing templates will be used to identify the corresponding fields in the NanoCommons KB. For more general databases, it needs to be determined whether curation templates and the resulting data schemas can be applied for all datasets in the database as a group, or whether they have to be handled on an individual dataset level. A similar approach will be applied for the required metadata and how they are handled, presented and made available. The list of ontologies used for data annotation will be required to test for compatibility and/or harmonisation with those used in NanoCommons (e.g. eNanoMapper and the extensions to this developed in NanoCommons, NanoSolveIT and ACEnano).
Details on data enrichment (processes of enhancing and refining raw data), if any, will be required. As Biomax is one of the leaders in the field of data enrichment internationally, such information will be needed to test for alignment and potential conflicts, and if not available can be offered to the source DB as an additional benefit of integration in NanoCommons.
Data governance and security
This section requires information on the security employed by the source DB to protect the submitted data and the entire infrastructure from malicious attacks. This includes the licensing system used and how that matches that used by NanoCommons, and will determine data FAIRness and accessibility to NanoCommons Users. GDPR compliance, ethics statements and links to ethical guidelines, ethical approval numbers and approving bodies will be requested where applicable.
Further information will be needed on the data quality tools implemented within the source DB. Such tools might include data profiling, cleansing, parsing and whether duplicate entries are easily or not identified or are systematically removed. The quality assurance processes used will be required to evaluate the quality distribution of the data transferred in response to NanoCommons User queries and the resulting big datasets.
Tools implementation
Finally, information on the implemented tools (e.g. searching and browsing, statistical analysis, predictive modelling, visualisation tools) along with any training materials will be required and their availability and accessibility via the KB assessed. All the information required from the source DBs in order to be integrated into NanoCommons are aligned with the principles of the unified Core Trustworthy Data Repository Requirements developed by the International Council for Science’s World Data System, which are also included in the source DB checklist. Additionally, NanoCommons infrastructure (KB / DW) also adheres to these principles, and is actively developing its long-term sustainability plan.
Appendix - Checklist for DB integration into the NanoCommons KB
- The repository provides detailed information on the OS, implemented tools and APIs used
- The repository provides the necessary information regarding its maximum load capacity and increased traffic handling
- The repository provides detailed mapping of its hierarchical structure to facilitate interoperability and synchronisation
- The repository provides information on the supported communication protocols for data movement, sharing and synchronisation
- The repository has put in place, and provides detailed information, a licensing system and appropriate security measures
- The repository maintains all applicable licenses covering data access and use and monitors compliance
- The repository has a continuity plan to ensure ongoing access to and preservation of its holdings
- The repository ensures, to the extent possible, that data are created, curated, accessed, and used in compliance with disciplinary and ethical norms
- The repository guarantees the integrity and authenticity of the data
- The repository accepts data and metadata based on defined criteria to ensure relevance and understandability for data users
- The repository assumes responsibility for long-term preservation and manages this function in a planned and documented way
- The repository has appropriate expertise to address technical data and metadata quality and ensures that sufficient information is available for end users to make quality-related evaluations
- The repository enables users to discover the data and refer to them in a persistent way through proper citation
- The repository enables reuse of the data over time, ensuring that appropriate metadata are available to support the understanding and use of the data